 
_tricks.sliding_window_view constructs views on numpyĪrrays that offer a sliding or moving window access to the array. ( gh-15121) sliding_window_view provides a sliding window view for numpy arrays # It is now possible to permute the rows or columns of a 2-D array. Subarrays indexed by an axis are permuted rather than the axis being treated asĪ separate 1-D array for every combination of the other indexes. The new function differs from shuffle and permutation in that the New functions # The random.Generator class has a new permuted function. Preliminary support for the upcoming Cython 3.0. 
Is ongoing and part of the larger project to improve NumPy’s online presenceįurther cleanups related to removing Python 2.7. Has been done to allow experimentation and feedback.Įxtensive documentation improvements comprising some 185 PR merges. Provide an easier path to extending dtypes. Preliminary work in changing the dtype and casting implementations in order to Much work has beenĭone in introducing universal functions that will ease use of modernįeatures across different hardware platforms. Wider use of SIMD to increase execution speed of ufuncs. This work is ongoing and improvements can The Python versions supported for this release are 3.7-3.9, support for PythonĪnnotations for NumPy functions. See the list of highlights below for more details. So you have an option of just displaying a leaf but have an option of showing (in parenthesis) how many are bunched up in that leaf.This NumPy release is the largest so made to date, some 684 PRs contributed byġ84 people have been merged. Basically a leaves are bunched up so close together that they are not easy to see. (In my example code above, all distances are Euclidean so all are positive and consistent from points on a 2d plane.)įor your second question, you probably need to roll out your own annotation routine to do what you want, since I don't think dendromgram natively supports it.įor the last question, show_leaf_counts seems to work only when you try to display non-singleton leaf nodes with truncate_mode='lastp' option. Labels=array(),ĭ.append(np.sqrt(((x-x)**2 + (y-y)**2)))įirst of all, the computation m -> m - 1 didn't really change your result since the distance matrix, which basically describes the relative distances between all unique pairs, didn't change in your specific case. Linkage_matrix = linkage(dist_mat, 'single') Here's a fully working code snippet to illustrate my points: import matplotlib.pyplot as pltįrom import dendrogram, linkage I think there's a couple misunderstandings as to the use of the functions that you are trying to use. The following shows the effect of show_leaf_counts. With p=6 and trunc_mode="lastp", dendrogram only shows the "top" Plt.title("show_leaf_counts = %s" % show_leaf_counts) X = np.random.multivariate_normal(, np.array(, ]),ĭdata = augmented_dendrogram(linkage_matrix, # Generate a random sample of `n` points in 2-d. Here's an example with 100 points: import numpy as npįrom import linkageįrom augmented_dendrogram import augmented_dendrogram The flag show_leaf_counts only applies when not all the original data So that the number of objects in each class is shown? It seems that show_leaf_counts flag is ignored, is there a way to turn it on So point 'a' and 'c' are 1.01 units apart, and point 'b' is 1.57 units from Plt.annotate("%.3g" % y, (x, y), xytext=(0, -8),įor your mat array, the augmented dendrogram is from import dendrogramĭef augmented_dendrogram(*args, **kwargs):įor i, d in zip(ddata, ddata): Is used to add a label of the distance (i.e. With the keys icoord and dcoord give the x and y coordinates of each Segments of the diagram with the corresponding distance. Show how you can use the data returned by dendrogram to label the horizontal How can I annotate the distance along each branch of the tree using dendrogram so that the distances between pairs of nodes can be compared? Nor a translation ( mat + offset) of the entire data set change the relative 
Is based on distances between the points, and neither a reflection ( -mat) The arrays mat and 1-mat produce the same clustering because the clustering Why does mat and 1-mat give identical clusterings here? 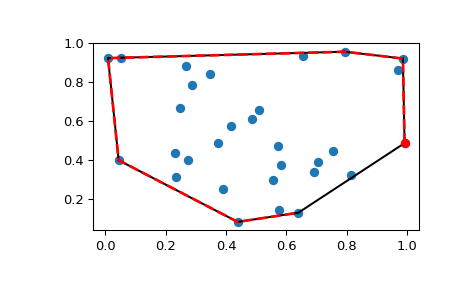
Clustering is based on the distance between these points. In your example, mat is 3 x 3, so you are clustering M-dimensional space, or a one-dimensional array containing the condensed distance matrix. The input to linkage() is either an n x m array, representing n points in 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
